CLM¶
Overview¶
~/cesm_?_?_? directory with the appropriate SourceMods structure. The
ensuing scripts require these SourceMods and expect them to be in your HOME directory.Script |
Description |
|---|---|
runs a single instance of CLM to harvest synthetic observations for an OSSE or “perfect model” experiment. It requires a single CLM state from a previous experiment and uses a specified DATM stream for forcing. This parallels an assimilation experiment in that in the multi-instance setting each CLM instance may use (should use?) a unique DATM forcing. This script has almost nothing to do with DART. There is one (trivial) section that records some configuration information in the DART setup script, but that’s about it. This script should initially be run without DART to ensure a working CESM environment. As of (V7195) 3 October 2014, this script demonstrates how to create ‘vector’-based CLM history files (which requires a bugfix) and has an option to use a bugfixed snow grain-size code. http://bugs.cgd.ucar.edu/show_bug.cgi?id=1730 http://bugs.cgd.ucar.edu/show_bug.cgi?id=1934 |
|
Is functionally identical to |
|
runs a multi-instance CLM experiment and can be used to perform a free run or ‘open loop’ experiment. By default, each CLM instance uses a unique DATM forcing. This script also has almost nothing to do with DART. There is one (trivial) section that records some configuration information in the DART setup script, but that’s about it. This script should initially be run without DART to ensure a working CESM. As of (V7195) 3 October 2014, this script demonstrates how to create ‘vector’-based CLM history files (which requires a bugfix) and has an option to use a bugfixed snow grain-size code. http://bugs.cgd.ucar.edu/show_bug.cgi?id=1730 http://bugs.cgd.ucar.edu/show_bug.cgi?id=1934 |
|
Is functionally identical to |
|
augments a CESM case with the bits and pieces required to
run DART. When either |
In addition to the script above, there are a couple scripts that will either perform an assimilation (assimilate.csh) or harvest observations for a perfect model experiment (perfect_model.csh). These scripts are designed to work on several compute platforms although they require configuration, mainly to indicate the location of the DART observation sequence files on your system.
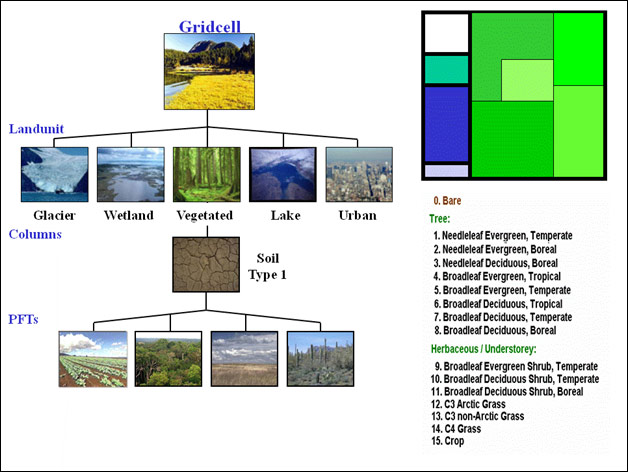
Pertinent details of the CLM gridcell¶
“The land surface is represented by 5 primary sub-grid land cover types (landunits: glacier, lake, wetland, urban, vegetated) in each grid cell. The vegetated portion of a grid cell is further divided into patches of plant functional types, each with its own leaf and stem area index and canopy height. Each subgrid land cover type and PFT patch is a separate column for energy and water calculations.” – CLM documentation. The only location information available is at the gridcell level. All landunits, columns, and PFTs in that gridcell have the same location. This has ramifications for the forward observation operators. If the observation metadata has information about land use/land cover, it can be used to select only those patches that are appropriate. Otherwise, an area-weighted average of ALL patches in the gridcell is used to calculate the observation value for that location. |
A word about forward observation operators¶
.h1.
history file. The smaller the CLM gridcell, the more likely it seems that these values will agree with point
observations..h1. history file needed for any particular time.Major changes as of (v7195) 3 october 2014¶
clm_vector_history_filename to support the concept that two history files can
be supported. My intent was to have the original history file (required for grid metadata) and another for support of
vector-based quantities in support of forward observation operators. Upon reflection, I’m not sure I need two
different history files - BUT - I’m sure there will be a situation where it comes in handy.input.nml model_nml demonstrates how to construct the DART state vector. The following table
explains in detail each entry for clm_variables:Column 1 |
Column 2 |
Column 3 |
Column 4 |
Column 5 |
Column 6 |
|---|---|---|---|---|---|
Variable name |
DART KIND |
minimum |
maximum |
filename |
update |
Column 1 |
Variable name |
This is the CLM variable name as it appears in the CLM netCDF file. |
Column 2 |
DART KIND |
This is the character string of the corresponding DART KIND. |
Column 3 |
minimum |
If the variable is to be updated in the CLM restart file, this specifies the minimum value. If set to ‘NA’, there is no minimum value. |
Column 4 |
maximum |
If the variable is to be updated in the CLM restart file, this specifies the maximum value. If set to ‘NA’, there is no maximum value. |
Column 5 |
filename |
This specifies which file should be
used to obtain the variable.
|
Column 6 |
update |
If the variable comes from the
restart file, it may be updated after
the assimilation.
|
The following are only meant to be examples - they are not scientifically validated. Some of these that are UPDATED are probably diagnostic quantities, Some of these that should be updated may be marked NO_COPY_BACK. There are multiple choices for some DART kinds. This list is by no means complete.
'livecrootc', 'QTY_ROOT_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'deadcrootc', 'QTY_ROOT_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'livestemc', 'QTY_STEM_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'deadstemc', 'QTY_STEM_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'livecrootn', 'QTY_ROOT_NITROGEN', 'NA', 'NA', 'restart', 'UPDATE',
'deadcrootn', 'QTY_ROOT_NITROGEN', 'NA', 'NA', 'restart', 'UPDATE',
'livestemn', 'QTY_STEM_NITROGEN', 'NA', 'NA', 'restart', 'UPDATE',
'deadstemn', 'QTY_STEM_NITROGEN', 'NA', 'NA', 'restart', 'UPDATE',
'litr1c', 'QTY_LEAF_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'litr2c', 'QTY_LEAF_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'litr3c', 'QTY_LEAF_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'soil1c', 'QTY_SOIL_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'soil2c', 'QTY_SOIL_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'soil3c', 'QTY_SOIL_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'soil4c', 'QTY_SOIL_CARBON', 'NA', 'NA', 'restart', 'UPDATE',
'fabd', 'QTY_FPAR_DIRECT', 'NA', 'NA', 'restart', 'UPDATE',
'fabi', 'QTY_FPAR_DIFFUSE', 'NA', 'NA', 'restart', 'UPDATE',
'T_VEG', 'QTY_VEGETATION_TEMPERATURE', 'NA', 'NA', 'restart', 'UPDATE',
'fabd_sun_z', 'QTY_FPAR_SUNLIT_DIRECT', 'NA', 'NA', 'restart', 'UPDATE',
'fabd_sha_z', 'QTY_FPAR_SUNLIT_DIFFUSE', 'NA', 'NA', 'restart', 'UPDATE',
'fabi_sun_z', 'QTY_FPAR_SHADED_DIRECT', 'NA', 'NA', 'restart', 'UPDATE',
'fabi_sha_z', 'QTY_FPAR_SHADED_DIFFUSE', 'NA', 'NA', 'restart', 'UPDATE',
'elai', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'restart', 'UPDATE',
Only the first variable for a DART kind in the clm_variables list will be used for the forward observation operator. The following is perfectly legal (for CLM4, at least):
clm_variables = 'LAIP_VALUE', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'restart' , 'UPDATE',
'tlai', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'restart' , 'UPDATE',
'elai', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'restart' , 'UPDATE',
'ELAI', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'history' , 'NO_COPY_BACK',
'LAISHA', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'history' , 'NO_COPY_BACK',
'LAISUN', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'history' , 'NO_COPY_BACK',
'TLAI', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'history' , 'NO_COPY_BACK',
'TLAI', 'QTY_LEAF_AREA_INDEX', 'NA', 'NA', 'vector' , 'NO_COPY_BACK'
/
however, only LAIP_VALUE will be used to calculate the LAI when an observation of LAI is encountered. All the other LAI variables in the DART state will be modified by the assimilation based on the relationship of LAIP_VALUE and the observation. Those coming from the restart file and marked ‘UPDATE’ will be updated in the CLM restart file.
Namelist¶
These namelists are read from the file input.nml. Namelists start with an ampersand ‘&’ and terminate with a slash
‘/’. Character strings that contain a ‘/’ must be enclosed in quotes to prevent them from prematurely terminating the
namelist.
&model_nml
clm_restart_filename = 'clm_restart.nc',
clm_history_filename = 'clm_history.nc',
clm_vector_history_filename = 'clm_vector_history.nc',
output_state_vector = .false.,
assimilation_period_days = 2,
assimilation_period_seconds = 0,
model_perturbation_amplitude = 0.2,
calendar = 'Gregorian',
debug = 0
clm_variables = 'frac_sno', 'QTY_SNOWCOVER_FRAC', 'NA' , 'NA', 'restart' , 'NO_COPY_BACK',
'H2OSNO', 'QTY_SNOW_WATER', '0.0', 'NA', 'restart' , 'UPDATE',
'H2OSOI_LIQ', 'QTY_SOIL_MOISTURE', '0.0', 'NA', 'restart' , 'UPDATE',
'H2OSOI_ICE', 'QTY_ICE', '0.0', 'NA', 'restart' , 'UPDATE',
'T_SOISNO', 'QTY_SOIL_TEMPERATURE', 'NA' , 'NA', 'restart' , 'UPDATE',
'SNOWDP', 'QTY_SNOW_THICKNESS', 'NA' , 'NA', 'restart' , 'UPDATE',
'LAIP_VALUE', 'QTY_LEAF_AREA_INDEX', 'NA' , 'NA', 'restart' , 'NO_COPY_BACK',
'cpool', 'QTY_CARBON', '0.0', 'NA', 'restart' , 'UPDATE',
'frootc', 'QTY_ROOT_CARBON', '0.0', 'NA', 'restart' , 'UPDATE',
'leafc', 'QTY_LEAF_CARBON', '0.0', 'NA', 'restart' , 'UPDATE',
'leafn', 'QTY_LEAF_NITROGEN', '0.0', 'NA', 'restart' , 'UPDATE',
'NEP', 'QTY_NET_CARBON_PRODUCTION', 'NA' , 'NA', 'history' , 'NO_COPY_BACK',
'TV', 'QTY_VEGETATION_TEMPERATURE', 'NA' , 'NA', 'vector' , 'NO_COPY_BACK',
'RH2M_R', 'QTY_SPECIFIC_HUMIDITY', 'NA' , 'NA', 'vector' , 'NO_COPY_BACK',
'PBOT', 'QTY_SURFACE_PRESSURE', 'NA' , 'NA', 'vector' , 'NO_COPY_BACK',
'TBOT', 'QTY_TEMPERATURE', 'NA' , 'NA', 'vector' , 'NO_COPY_BACK'
/
Item |
Type |
Description |
|---|---|---|
clm_restart_filename |
character(len=256) |
this is the filename of the CLM restart file. The DART scripts resolve linking the specific CLM restart file to this generic name. This file provides the elements used to make up the DART state vector. The variables are in their original landunit, column, and PFT-based representations. |
clm_history_filename |
character(len=256) |
this is the filename of the CLM
|
clm_vector_history_filename |
character(len=256) |
this is the filename of a second CLM
history file. The DART scripts
resolve linking the specific CLM
history file to this generic name.
The default setup scripts actually
create 3 separate CLM history files,
the |
output_state_vector |
logical |
If .true. write state vector as a 1D array to the DART diagnostic output files. If .false. break state vector up into variables before writing to the output files. |
assimilation_period_days, assimilation_period_seconds |
integer |
Combined, these specify the width of the assimilation window. The current model time is used as the center time of the assimilation window. All observations in the assimilation window are assimilated. BEWARE: if you put observations that occur before the beginning of the assimilation_period, DART will error out because it cannot move the model ‘back in time’ to process these observations. |
model_perturbation_amplitude |
real(r8) |
Required by the DART interfaces, but not used by CLM. |
calendar |
character(len=32) |
string specifying the calendar to use with DART. The CLM dates will be interpreted with this same calendar. For assimilations with real observations, this should be ‘Gregorian’. |
debug |
integer |
Set to 0 (zero) for minimal output. Successively higher values generate successively more output. Not all values are important, however. It seems I’ve only used values [3,6,7,8]. Go figure. |
clm_state_variables clm_variables |
character(:,6) |
Strings that identify the CLM
variables, their DART kind, the min &
max values, what file to read from,
and whether or not the file should be
updated after the assimilation. Only
CLM variable names in the CLM restart
file are valid. The DART kind must
be one found in the
|
&obs_def_tower_nml
casename = '../clm_dart',
hist_nhtfrq = -24,
debug = .false.
/
Item |
Type |
Description |
|---|---|---|
casename |
character(len=256) |
this is the name of the CESM case. It is used by the forward observation
operators to help construct the filename of the CLM |
hist_nhtfrq |
integer |
this is the same value as in the CLM documentation. A negative value indicates
the number of hours contained in the |
debug |
logical |
Set to .false. for minimal output. |
Other modules used (directly)¶
types_mod
time_manager_mod
threed_sphere/location_mod
utilities_mod
obs_kind_mod
obs_def_land_mod
obs_def_tower_mod
random_seq_mod
Public interfaces - required¶
use model_mod, only : |
get_model_size |
adv_1step |
|
get_state_meta_data |
|
model_interpolate |
|
get_model_time_step |
|
static_init_model |
|
end_model |
|
init_time |
|
init_conditions |
|
nc_write_model_atts |
|
nc_write_model_vars |
|
pert_model_state |
|
get_close_maxdist_init |
|
get_close_obs_init |
|
get_close_obs |
|
ens_mean_for_model |
A note about documentation style. Optional arguments are enclosed in brackets [like this].
model_size = get_model_size( )
integer :: get_model_size
Returns the length of the model state vector.
|
The length of the model state vector. |
call adv_1step(x, time)
real(r8), dimension(:), intent(inout) :: x
type(time_type), intent(in) :: time
Advances the model for a single time step. The time associated with the initial model state is also input although it is not used for the computation.
|
State vector of length model_size. |
|
Specifies time of the initial model state. |
call get_state_meta_data (index_in, location, [, var_type] )
integer, intent(in) :: index_in
type(location_type), intent(out) :: location
integer, optional, intent(out) :: var_type
Returns metadata about a given element, indexed by index_in, in the model state vector. The location defines where the state variable is located.
|
Index of state vector element about which information is requested. |
|
The location of state variable element. |
var_type |
The generic DART kind of the state variable element. |
call model_interpolate(x, location, itype, obs_val, istatus)
real(r8), dimension(:), intent(in) :: x
type(location_type), intent(in) :: location
integer, intent(in) :: itype
real(r8), intent(out) :: obs_val
integer, intent(out) :: istatus
Given model state, returns the value interpolated to a given location.
|
A model state vector. |
|
Location to which to interpolate. |
|
Not used. |
|
The interpolated value from the model. |
|
If the interpolation was successful |
var = get_model_time_step()
type(time_type) :: get_model_time_step
Returns the time step (forecast length) of the model;
|
Smallest time step of model. |
call static_init_model()
Used for runtime initialization of model; reads namelist, initializes model parameters, etc. This is the first call made to the model by any DART-compliant assimilation routine.
call end_model()
A stub.
call init_time(time)
type(time_type), intent(out) :: time
Returns the time at which the model will start if no input initial conditions are to be used. This is used to spin-up the model from rest.
|
Initial model time. |
call init_conditions(x)
real(r8), dimension(:), intent(out) :: x
Returns default initial conditions for the model; generally used for spinning up initial model states.
|
Initial conditions for state vector. |
ierr = nc_write_model_atts(ncFileID)
integer :: nc_write_model_atts
integer, intent(in) :: ncFileID
Function to write model specific attributes to a netCDF file. At present, DART is using the NetCDF format to output diagnostic information. This is not a requirement, and models could choose to provide output in other formats. This function writes the metadata associated with the model to a NetCDF file opened to a file identified by ncFileID.
|
Integer file descriptor to previously-opened netCDF file. |
|
Returns a 0 for successful completion. |
ierr = nc_write_model_vars(ncFileID, statevec, copyindex, timeindex)
integer :: nc_write_model_vars
integer, intent(in) :: ncFileID
real(r8), dimension(:), intent(in) :: statevec
integer, intent(in) :: copyindex
integer, intent(in) :: timeindex
Writes a copy of the state variables to a netCDF file. Multiple copies of the state for a given time are supported, allowing, for instance, a single file to include multiple ensemble estimates of the state.
|
file descriptor to previously-opened netCDF file. |
|
A model state vector. |
|
Integer index of copy to be written. |
|
The timestep counter for the given state. |
|
Returns 0 for normal completion. |
call pert_model_state(state, pert_state, interf_provided)
real(r8), dimension(:), intent(in) :: state
real(r8), dimension(:), intent(out) :: pert_state
logical, intent(out) :: interf_provided
Given a model state, produces a perturbed model state.
|
State vector to be perturbed. |
|
Perturbed state vector: NOT returned. |
|
Returned false; interface is not implemented. |
call get_close_maxdist_init(gc, maxdist)
type(get_close_type), intent(inout) :: gc
real(r8), intent(in) :: maxdist
In distance computations any two locations closer than the given maxdist will be considered close by the
get_close_obs() routine. Pass-through to the 3D Sphere locations module. See
get_close_maxdist_init()
for the documentation of this subroutine.
call get_close_obs_init(gc, num, obs)
type(get_close_type), intent(inout) :: gc
integer, intent(in) :: num
type(location_type), intent(in) :: obs(num)
Pass-through to the 3D Sphere locations module. See get_close_obs_init() for the documentation of this subroutine.
call get_close_obs(gc, base_obs_loc, base_obs_kind, obs, obs_kind, num_close, close_ind [, dist])
type(get_close_type), intent(in) :: gc
type(location_type), intent(in) :: base_obs_loc
integer, intent(in) :: base_obs_kind
type(location_type), intent(in) :: obs(:)
integer, intent(in) :: obs_kind(:)
integer, intent(out) :: num_close
integer, intent(out) :: close_ind(:)
real(r8), optional, intent(out) :: dist(:)
Pass-through to the 3D Sphere locations module. See get_close_obs() for the documentation of this subroutine.
call ens_mean_for_model(ens_mean)
real(r8), dimension(:), intent(in) :: ens_mean
A NULL INTERFACE in this model.
|
State vector containing the ensemble mean. |
Public interfaces - optional¶
use model_mod, only : |
get_gridsize |
clm_to_dart_state_vector |
|
sv_to_restart_file |
|
get_clm_restart_filename |
|
get_state_time |
|
get_grid_vertval |
|
compute_gridcell_value |
|
gridcell_components |
|
DART_get_var |
|
get_model_time |
call get_gridsize(num_lon, num_lat, num_lev)
integer, intent(out) :: num_lon, num_lat, num_lev
Returns the number of longitudes, latitudes, and total number of levels in the CLM state.
|
The number of longitude grid cells in the CLM state. This comes from the CLM history file. |
|
The number of latitude grid cells in the CLM state. This comes from the CLM history file. |
|
The number of levels grid cells in the CLM state. This comes from ‘nlevtot’ in the CLM restart file. |
call clm_to_dart_state_vector(state_vector, restart_time)
real(r8), intent(inout) :: state_vector(:)
type(time_type), intent(out) :: restart_time
model_nml:clm_variables. Each variable specifies its own file of origin. If there are multiple times in the
file of origin, only the time that matches the restart file are used.
|
The DART state vector. |
|
The valid time of the CLM state. |
call sv_to_restart_file(state_vector, filename, dart_time)
real(r8), intent(in) :: state_vector(:)
character(len=*), intent(in) :: filename
type(time_type), intent(in) :: dart_time
This routine updates the CLM restart file with the posterior state from the assimilation. Some CLM variables that are
useful to include in the DART state (frac_sno, for example) are diagnostic quantities and are not used for subsequent
model advances. The known diagnostic variables are NOT updated. If the values created by the assimilation are outside
physical bounds, or if the original CLM value was ‘missing’, the vector_to_prog_var() subroutine ensures that the
values in the original CLM restart file are not updated.
|
The DART state vector containing the state modified by the assimilation. |
|
The name of the CLM restart file. The contents of some of the variables will be overwritten with new values. |
|
The valid time of the DART state. This has to match the time in the CLM restart file. |
call get_clm_restart_filename( filename )
character(len=*), intent(out) :: filename
provides access to the name of the CLM restart file to routines outside the scope of this module.
|
The name of the CLM restart file. |
time = get_state_time(file_handle)
integer, intent(in) :: file_handle
character(len=*), intent(in) :: file_handle
type(time_type) :: get_state_time
This routine has two interfaces - one for an integer input, one for a filename. They both return the valid time of the model state contained in the file. The file referenced is the CLM restart file in netCDF format.
|
If specified as an integer, it must be the netCDF file identifier from nf90_open(). If specified as a filename, the name of the netCDF file. |
|
A DART time-type that contains the valid time of the model state in the CLM restart file. |
call get_grid_vertval(x, location, varstring, interp_val, istatus)
real(r8), intent(in) :: x(:)
type(location_type), intent(in) :: location
character(len=*), intent(in) :: varstring
real(r8), intent(out) :: interp_val
integer, intent(out) :: istatus
Calculate the value of quantity at depth. The gridcell value at the levels above and below the depth of interest are
calculated and then the value for the desired depth is linearly interpolated. Each gridcell value is an area-weighted
value of an unknown number of column- or pft-based quantities. This is one of the workhorse routines for
model_interpolate().
|
The DART state vector. |
|
The location of the desired quantity. |
|
The CLM variable of interest - this must be part of the DART state. e.g, T_SOISNO, H2OSOI_LIQ, H2OSOI_ICE … |
|
The quantity at the location of interest. |
|
error code. 0 (zero) indicates a successful interpolation. |
call compute_gridcell_value(x, location, varstring, interp_val, istatus)
real(r8), intent(in) :: x(:)
type(location_type), intent(in) :: location
character(len=*), intent(in) :: varstring
real(r8), intent(out) :: interp_val
integer, intent(out) :: istatus
Calculate the value of a CLM variable in the DART state vector given a location. Since the CLM location information
is only available at the gridcell level, all the columns in a gridcell are area-weighted to derive the value for the
location. This is one of the workhorse routines for model_interpolate(), and only select CLM variables are
currently supported. Only CLM variables that have no vertical levels may use this routine.
|
The DART state vector. |
|
The location of the desired quantity. |
|
The CLM variable of interest - this must be part of the DART state. e.g, frac_sno, leafc, ZWT … |
|
The quantity at the location of interest. |
|
error code. 0 (zero) indicates a successful interpolation. |
call gridcell_components(varstring)
character(len=*), intent(in) :: varstring
This is a utility routine that helps identify how many land units,columns, or PFTs are in each gridcell for a particular variable. Helps answer exploratory questions about which gridcells are appropriate to test code. The CLM variable is read from the CLM restart file.
|
The CLM variable name of interest. |
call DART_get_var(ncid, varname, datmat)
integer, intent(in) :: ncid
character(len=*), intent(in) :: varname
real(r8), dimension(:), intent(out) :: datmat
real(r8), dimension(:,:), intent(out) :: datmat
Reads a 1D or 2D variable of ‘any’ type from a netCDF file and processes and applies the offset/scale/FillValue attributes correctly.
|
The netCDF file identifier to an open file. ncid is the output from a nf90_open() call. |
|
The name of the netCDF variable of interest. The variables can be integers, floats, or doubles. |
|
The shape of datmat must match the shape of the netCDF variable. Only 1D or 2D variables are currently supported. |
model_time = get_model_time( )
integer :: get_model_time
Returns the valid time of the model state vector.
|
The valid time of the model state vector. |
Files¶
filename |
purpose |
|---|---|
input.nml |
to read the model_mod namelist |
clm_restart.nc |
both read and modified by the CLM model_mod |
clm_history.nc |
read by the CLM model_mod for metadata purposes. |
*.h1.* history files |
may be read by the obs_def_tower_mod for observation operator purposes. |
dart_log.out |
the run-time diagnostic output |
dart_log.nml |
the record of all the namelists actually USED - contains the default values |
References¶
CLM User’s Guide is an excellent reference for CLM.
Private components¶
N/A